Research
Quantitative genetics
To what extent can we predict quantitative phenotypes based on genotype? Mutations affect phenotype by perturbing the underlying cellular network: they can alter 1) enzymatic quality of proteins, 2) gene expression, and 3) protein folding amongst others. Focusing on the genetic basis of traits, we are broadly interested in asking what are the molecular processes that determine differences between individuals and therefore are acted on by evolutionary forces.
Systems biology of complex traits
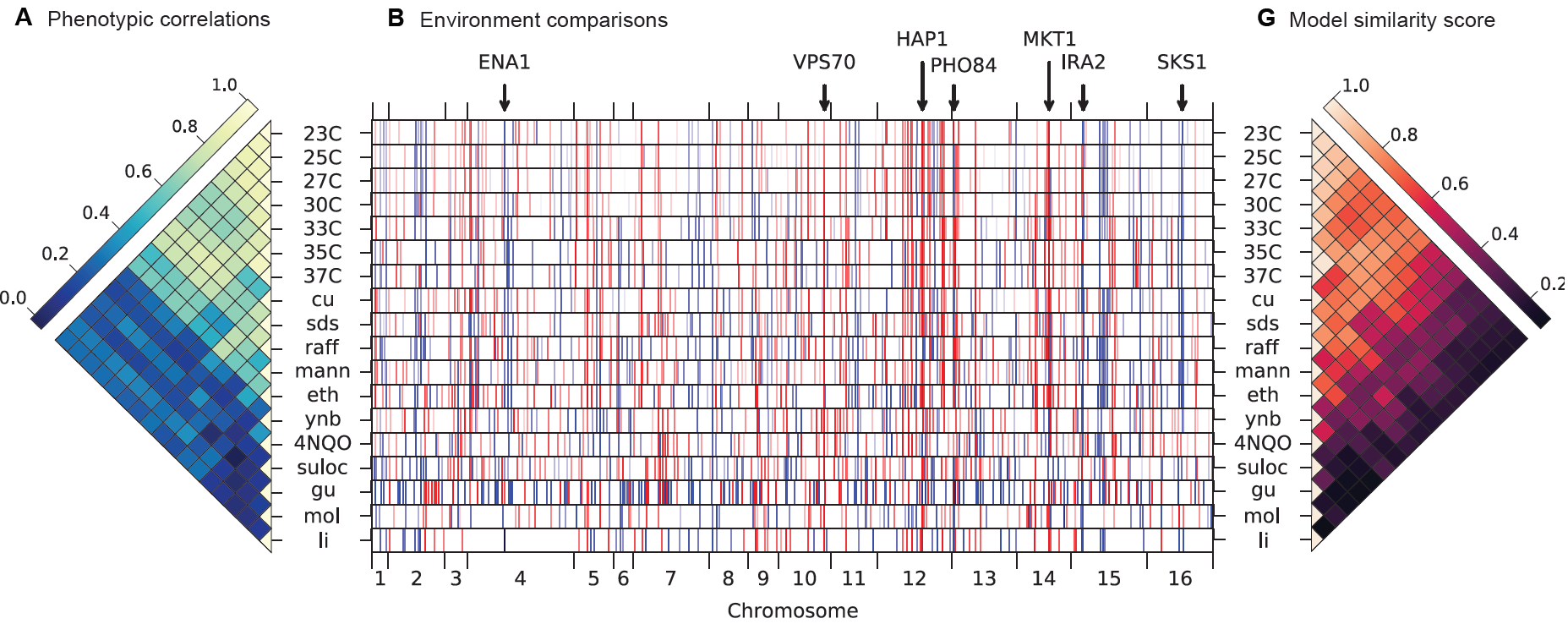
Thousands of polymorphisms in populations are responsible for phenotypic differences between individuals and between species. Which 'molecular knobs' are affected and are the main players between phenotypic divergence? Our research uses extremely large panels of derived strains from the wild to get at the systems biology of the cell.
Precision medicine
Our lab is participating in the effort to systematically characterize all functional missense variation in human genes. Our goal is to determine disease risks such as cancer before manifestation of symptoms (shifting medicine to a preventive science), and to reveal genetic determinants of inter-personal differences in molecular pathology to design better treatments,
Relevance
We hope to develop machine learning models of the cell to bring new insight into biological diseases. Examples of current projects aim to develop technologies for regenerative medicine in addressing aggregation-prone proteins found in Parkinson's and for screening disregulations that lead to cancers.
Evolution
Comparative genomics is the most popular approach to studying evolution. In contrast, our approach is to watch evolution as it happens, in the context of experiments that are performed under controlled conditions. We are interested in the dynamical process of selection and the evolutionary forces behind phenotypic changes. Ultimately, we hope to leverage our understanding of these processes to predict evolution.
Phenotypic evolution
Our lab makes use of several experimental evolution experiments to dissect the cellular basis of adaptation. For example, we know that transcriptional regulation plays a major role in cellular differentiation. However, how evolution tinkers with regulatory networks is less clear.
Vaccinology
B-cells undergo a selection process in germinal centers akin to evolutionary dynamics in microbial populations. A quantitative modeling of these processes is critical for developing an understanding of the repeatability of this evolutionary system and how it can be applied to vaccination success.
Our lab aims to understand how vaccine design is related to the underlying evolutionary process behind protective B-cell lineages.
Relevance
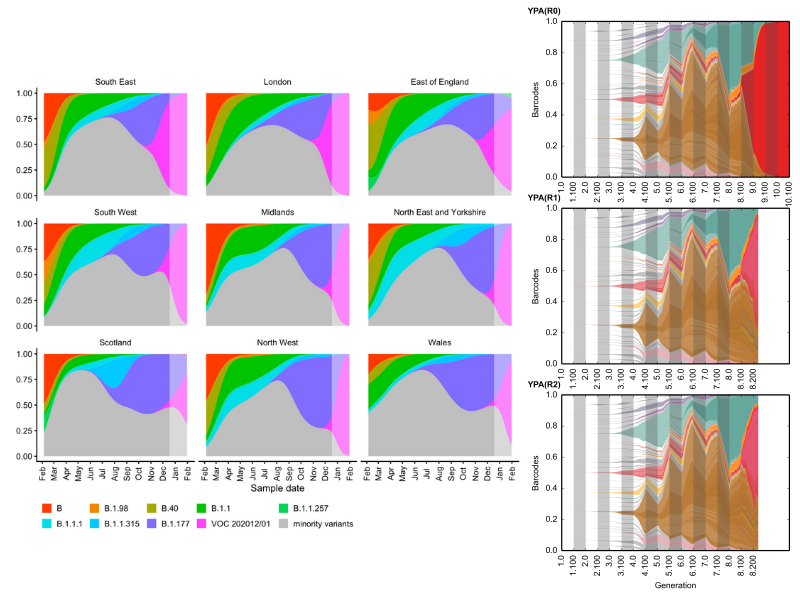
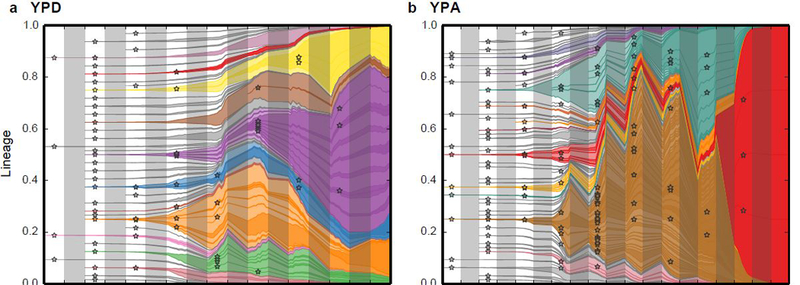
Our work in laboratory environments allows us to determine the relevant parameters of evolution, which can help in understanding and forecasting evolution in the real world. For example, note the similarity between the evolution of the Sars-CoV-2 variants of concern from Davies et al, 2020 (left), when compared to high-resolution lineage tracking of yeast populations in a flask (repeated evolution; right) :